Overview
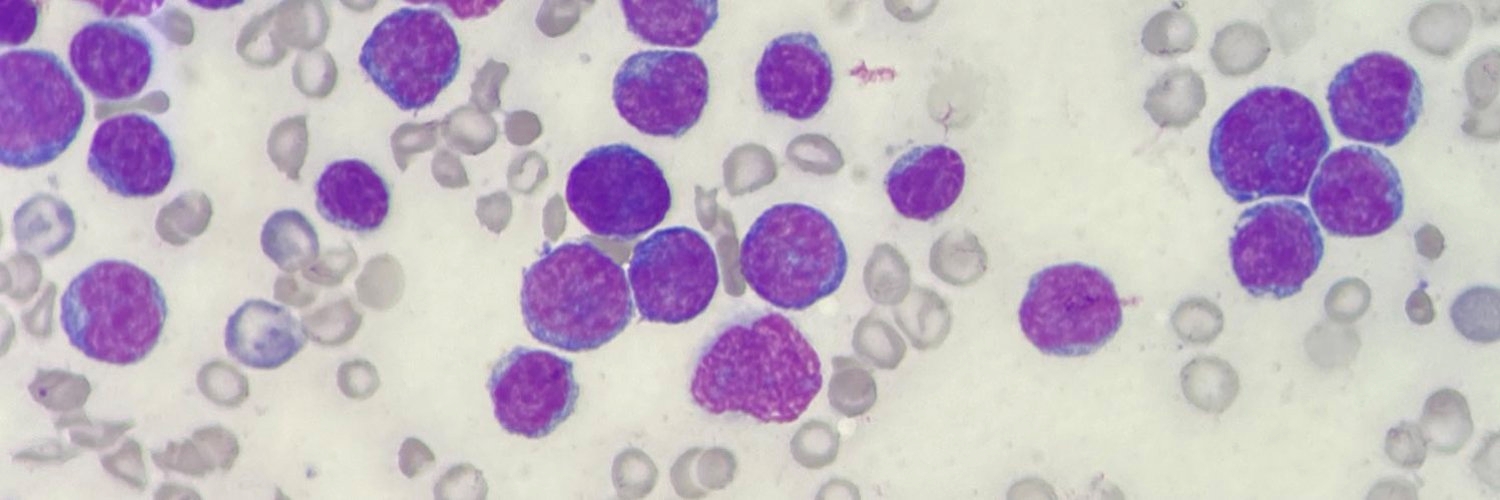
We study a blood cancer treatment called allogeneic haematopoietic stem cell transplantation. This is a complex therapy that involves treating patients with chemo/radiotherapy before infusing blood-forming stem cells harvested from a donor. Donor immune cells then eliminate residual leukaemia, a process termed the graft-versus-leukaemia effect. Stem cell transplantation can cure patients with blood cancers that would recur if managed with chemotherapy alone, but disease relapse is common and remains the leading cause of death. We believe that a better understanding of how this treatment works and why it fails will improve transplant outcomes and inform the development of novel immunotherapies for blood cancers.
Our group aims to understand and manipulate the graft-versus-leukaemia effect to reduce the risk of relapse and improve transplant outcomes. We also study fundamental questions of leukaemia immunology as these are critical to understanding how leukaemia cells interact with T cells and what determines the outcomes of these interactions.
We are studying the T cells found in bone marrow at leukaemia diagnosis and following stem cell transplantation. We have used mass cytometry to profile millions of T cells from hundreds of patients and are currently determining the ability of various leukaemia-associated populations to recognise cancer cells. At disease presentation we aim to establish whether autologous T-cell responses to AML are common and robust enough to be manipulated for therapeutic effect. At post-transplant relapse we aim to determine why leukaemia-reactive T cells lose control of the disease by identifying drivers of T-cell dysfunction.
AML is characterised by a block to differentiation leading to the accumulation of cancer cells that resemble myeloid stem and progenitor populations. Many recent and novel AML treatments work by releasing this block, leading to differentiation of leukaemia into mature myeloid cells, some of which are capable of presenting antigen to T helper cells. We are therefore investigating whether inducing leukaemic differentiation can support T-cell responses to clear the disease. Finally, we are developing novel biomarkers to predict the most serious transplant complications, namely disease relapse and graft-versus-host disease.
Featured Publications
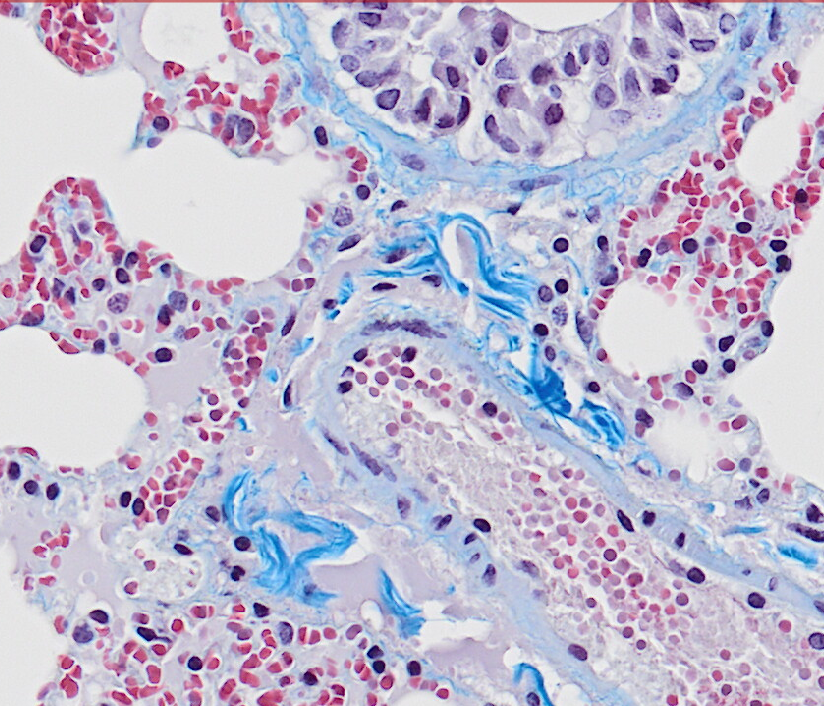
Targeted nanopore sequencing for the identification of ABCB1 promoter translocations in cancer.
10th November 2020
Resistance to chemotherapy is the most common cause of treatment failure in AML and drug efflux pump ABCB1 is a critical mediator. We established an experimental system and analysis pipeline to determine whether promoter translocations account for high ABCB1 expression in cases of relapsed human AML.
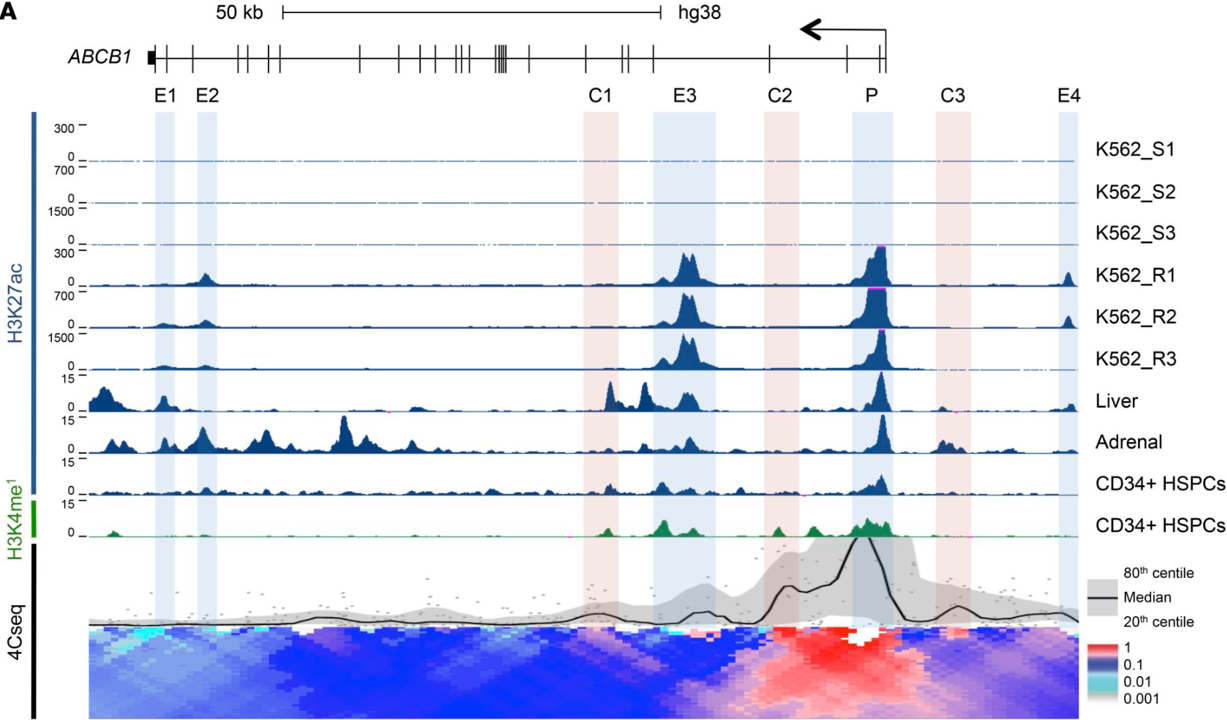
A stress-responsive enhancer induces dynamic drug resistance in acute myeloid leukemia.
4th February 2020
The drug efflux pump ABCB1 is a key driver of chemoresistance, and high expression predicts treatment failure in acute myeloid leukemia (AML). In this study, we identified and functionally validated the network of enhancers that controls expression of ABCB1.
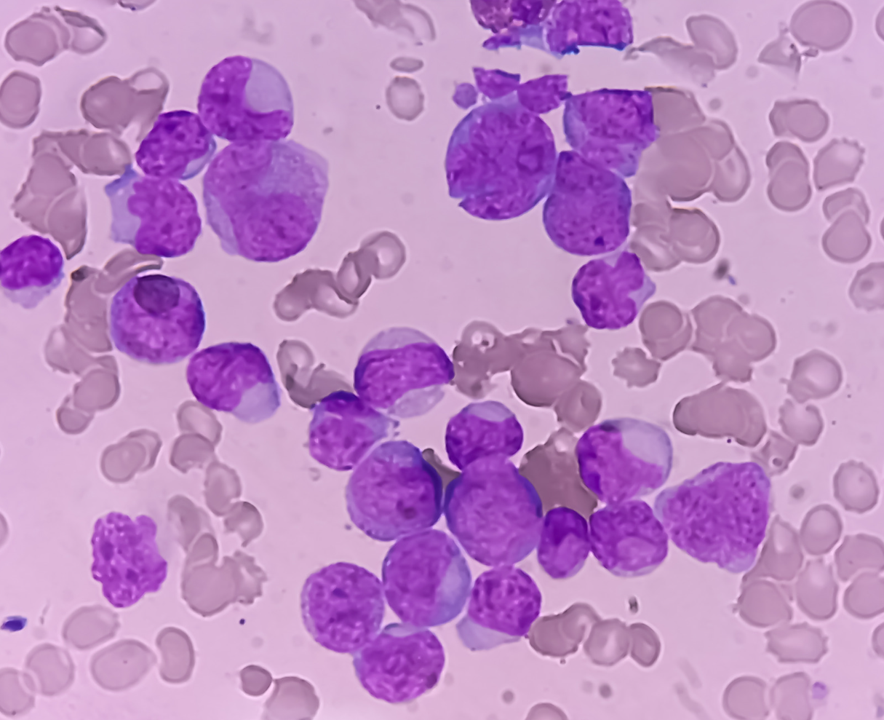
Blast cells surviving acute myeloid leukemia induction therapy are in cycle with a signature of FOXM1 activity
28th October 2021
Disease relapse remains common following treatment of acute myeloid leukemia due to chemoresistance of leukemia cells with disease repopulating potential. This paper shows chemorefractory blasts from leukemias with varied genetic backgrounds expressed a common transcriptional programme and the refractory blast gene signature implicates the transcription factor FOXM1.
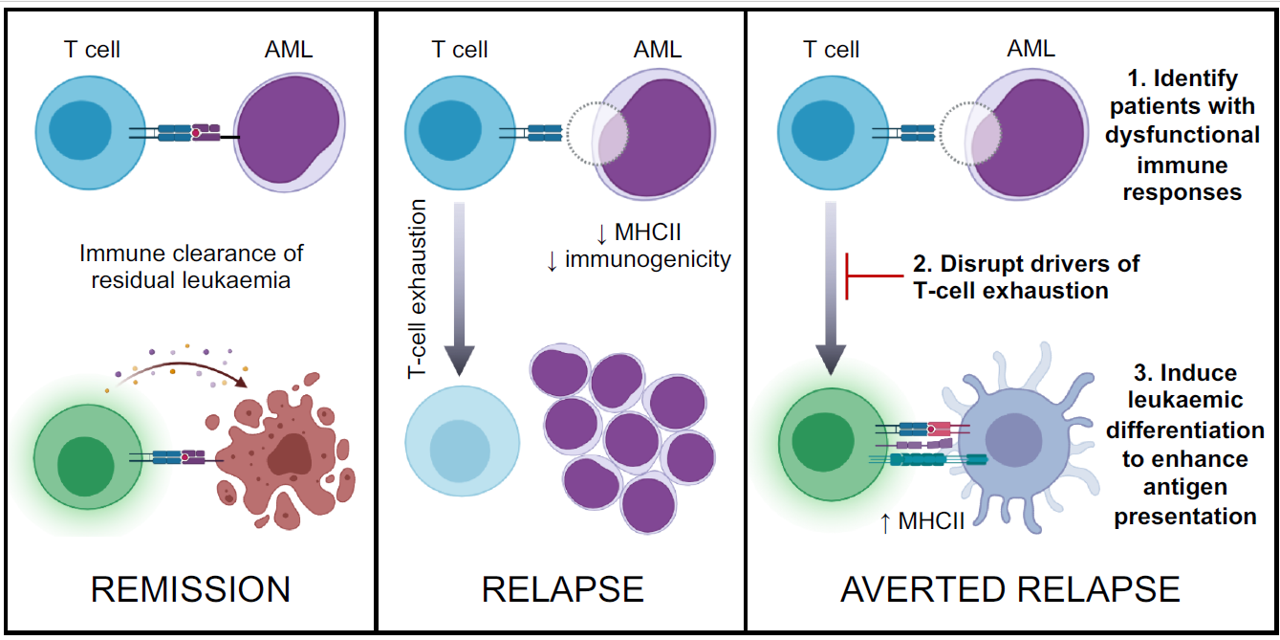
Strategy for preventing post-transplant relapse
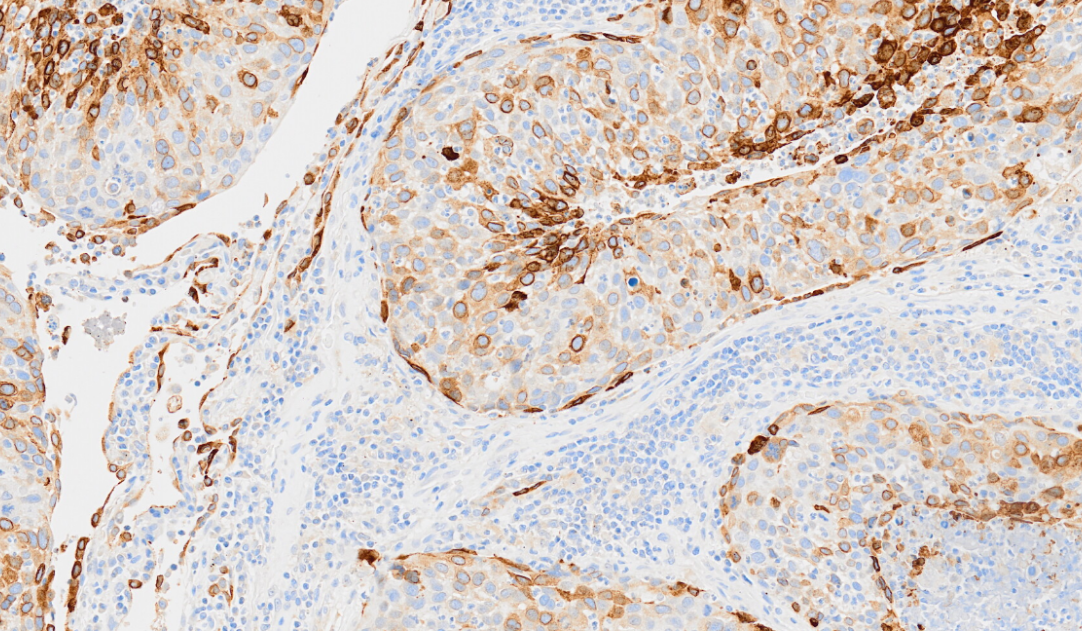
A strategy for preventing post-transplant AML relapse by identifying patients with early evidence of immune dysfunction then intervening to modify both T cell- and AML-mediated mechanisms of relapse, namely T-cell exhaustion and leukaemic immune evasion through downregulation of MHC class II (MHCII).
Meet the group
I am delighted to introduce my excellent team who work tirelessly on complex problems in order to improve the care of patients with blood cancers.

Institute Fellow
FAQs
Yes, I continue to see NHS patients in my weekly transplant clinic, it is their experience that informs and motivates the work we do. The Manchester Cancer Research Centre’s Biobank has built a large collection of blood and bone marrow samples from patients who kindly agree to donate. Much of our experimental work is done using these samples. We are also involved in developing clinical trials.
We work with research partners across the University of Manchester, around the UK and internationally. Get in touch with us using the contact form, we’re always open to new ideas and collaborations.
We are not advertising for PhD students currently, but please monitor the student pages [link] for details of upcoming studentship opportunities.
All Institute Publications
https://www.nature.com/articles/s42255-026-01514-y
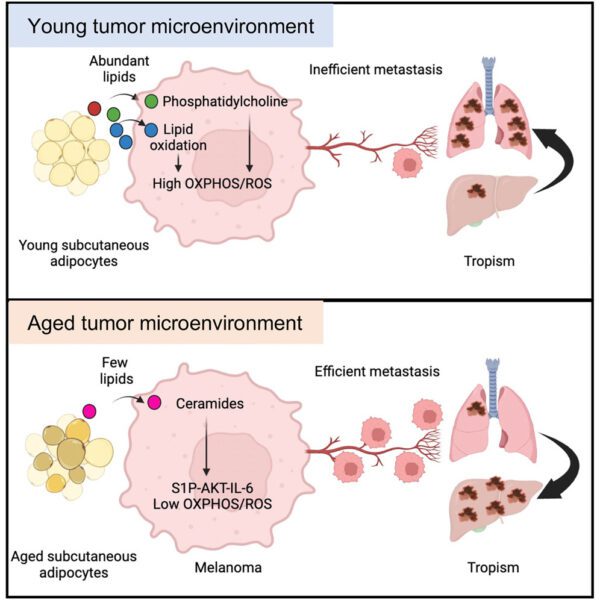
Tissue-specific fibroblast lipid cues impose the rate of epithelial cancer invasion
27 April 2026
Institute Authors (4)
Amaya Virós, Noah Palombo, Charlotte Russell, Claus Jørgensen
Research Group
Skin Cancer & Ageing
27 April 2026
https://www.cell.com/cancer-cell/fulltext/S1535-6108(26)00114-5
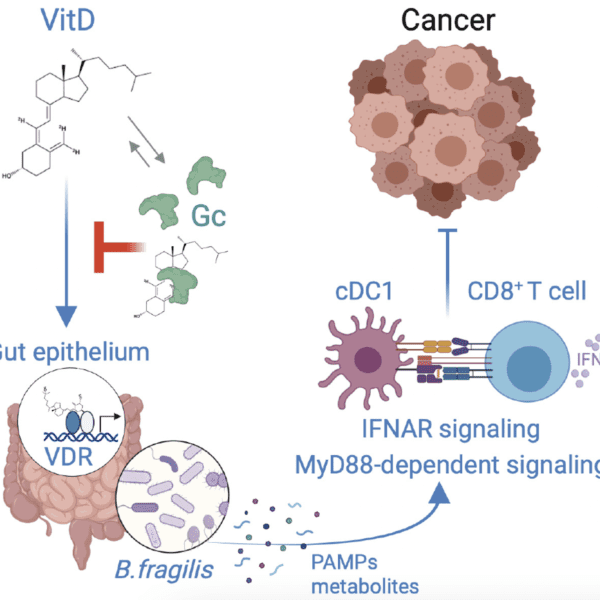
Immunometabolic gatekeeping: How tissue metabolism conditions tumor immunity
13 April 2026
Institute Authors (1)
Samra Turajlić
Research Group
Cancer Dynamics
13 April 2026
https://www.nature.com/articles/s41467-026-69964-2
Disruption of tRNA threonylation triggers RIG-I mediated anti-tumour immune response
25 February 2026
Institute Authors (1)
Sylvain Delaunay
Research Group
RNA Dynamics in Cancer
25 February 2026
https://doi.org/10.1038/s44161-025-00740-z
Single-cell profiling reveals three endothelial-to-hematopoietic transitions with divergent isoform expression landscapes
11 November 2025
Institute Authors (6)
Robert Sellers, John Weightman, Wolfgang Breitwieser, Natalia Moncaut, Michael Lie-a-ling, Georges Lacaud
Labs & Facilities
Computational Biology Support, Molecular Biology, Genome Editing and Mouse Models
Research Group
Stem Cell Biology
11 November 2025
https://doi.org/10.1136/jitc-2025-012527
Systemic immunosuppression from ultraviolet radiation exposure inhibits cancer immunotherapy
31 October 2025
Institute Authors (4)
Isabella Mataloni, Antonia Banyard, Garry Ashton, Amaya Virós
Labs & Facilities
Mass and Flow Cytometry, Histology
Research Group
Skin Cancer & Ageing
31 October 2025
https://aacrjournals.org/cancerdiscovery/article/doi/10.1158/2159-8290.CD-24-1224/766638/Glucocorticoids-Unleash-Immune-dependent-Melanoma
Glucocorticoids Unleash Immune-dependent Melanoma Control through Inhibition of the GARP/TGF β Axis
15 October 2025
Institute Authors (12)
Charles Earnshaw, Poppy Dunn, Shih-Chieh Chiang, Maria Koufaki, Massimo Russo, Kimberley Hockenhull, Erin Richardson, Anna Pidoux, Alex Baker, Richard Reeves, Robert Sellers, Sudhakar Sahoo
Labs & Facilities
Computational Biology Support, Visualisation, Irradiation and Analysis
Research Group
Cancer Inflammation and Immunity
15 October 2025
Get in touch
Our vision for world leading cancer research in the heart of Manchester
We are a leading cancer research institute within The University of Manchester, spanning the whole spectrum of cancer research – from investigating the molecular and cellular basis of cancer, to translational research and the development of therapeutics.
Our collaborations
Bringing together internationally renowned scientists and clinicians
Scientific Advisory Board
Supported by an international Scientific Advisory Board
Careers that have a lasting impact on cancer research and patient care
We are always on the lookout for talented and motivated people to join us. Whether your background is in biological or chemical sciences, mathematics or finance, computer science or logistics, use the links below to see roles across the Institute in our core facilities, operations teams, research groups, and studentships within our exceptional graduate programme.










A note from the Group Leader – Mark Williams
My research is focused on using the immune system to eliminate blood cancers. I am particularly keen to understand stem cell transplantation, as this remains the only approach that provides durable remission for patients with poor-risk haematological malignancies. Despite decades of clinical experience, we still don’t fully understand how transplant works. We are therefore unable to predict who will be cured and who will relapse. We also lack means to increase the potency of the therapeutic effect. There is tremendous potential to reimagine transplantation to make it a more effective and safer treatment, if only we could better understand the curative mechanism.