Overview
We are particularly interested in the chemical messages that are sent between cells as pancreatic cancer develops. We aim to identify the key signals that drive tumour growth and help the cancer cells to spread around the body, and ultimately find new ways to block these signals – reducing tumour growth and metastatic spread. Survival from pancreatic cancer has changed little over the years; the average five-year survival rate is just 12%, thus identifying new ways to treat this disease is a real priority for the 10,000 people diagnosed each year in the UK.
Pancreatic Ductal Adenocarcinoma is the most common form of Pancreatic Cancer. Its commonly detected all too late and most patients are presenting with metastatic disease. Local disease e.g., tumour that has not escaped the pancreas, may be removed by surgery and treated by chemotherapy whereas metastatic disease is only treated by chemotherapy. Pancreatic Cancer is characterised by an extensive desmoplastic reaction where most of the tumour is made up by host ‘normal’ cells and extracellular matrix (scar tissue) with tumour cells only occupying ~15% of the tumour volume on average. Host cells such as fibroblasts and various immune cell populations are co-opted by tumour cells to enable tumour development, resist treatment and avoid recognition by tumour cell killing immune cells. The remodelled extracellular matrix increases tissue stiffness and interstitial pressure, decreasing therapeutic efficiency, and altering cell growth. Our long-term goals are to understand how specific signals are processed to promote tumour progression and how blocking these signals can enhance therapeutic response.
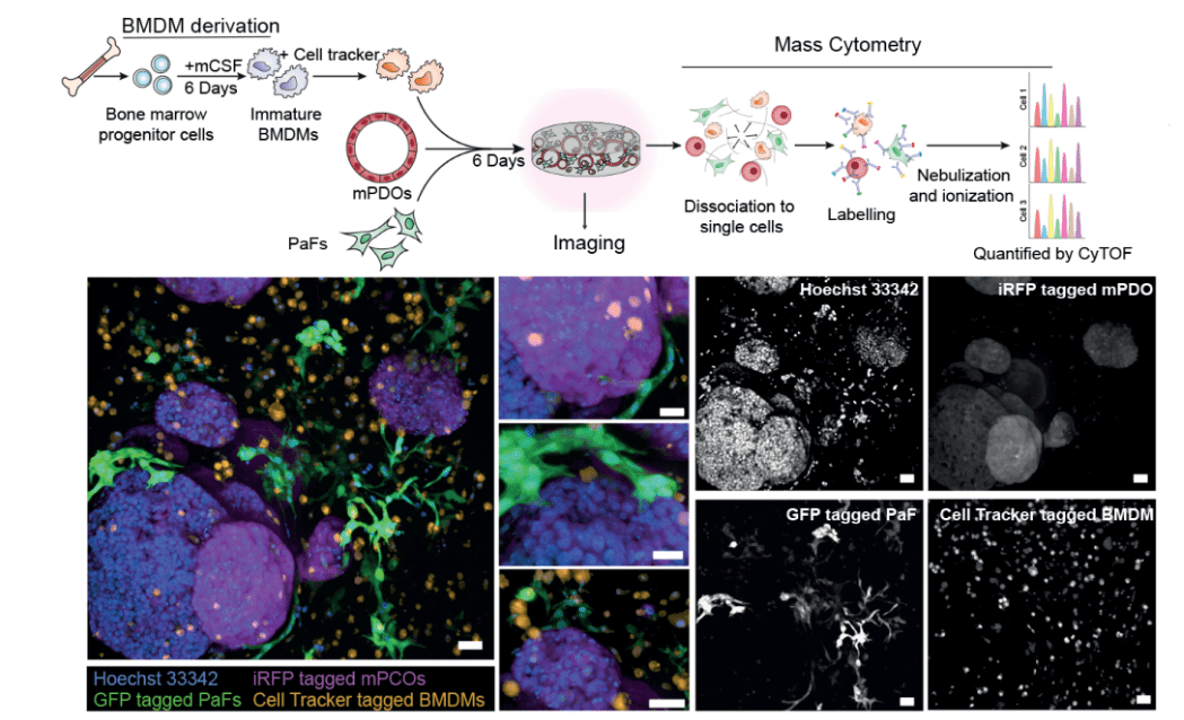
We focus on identifying and delineating the regulatory mechanisms whereby tumour cells co-opt stromal cells and how the ‘activated’ stromal cells affect tumour cell behaviour (malignant progression, drug sensitivity and metastasis). To establish the specificities of signalling networks and how they impinge on specific cellular processes, we are using Mass Cytometry (CyTOF), CRISPR editing/screening, sequencing and quantitative mass spectrometry to systematically measure signalling on a global level. Through data-integrative approaches, we aim to describe how signals are processed in a cell-specific manner, which requires single cell methodologies to separate the activities of all the cell types in the tumour. We are also creating new models of pancreatic cancer in the laboratory, using synthetic scaffolds to artificially alter the stiffness of the tumour, mixing in fibroblasts and immune cells, and experimentally determining the effect of each component on the tumour growth.
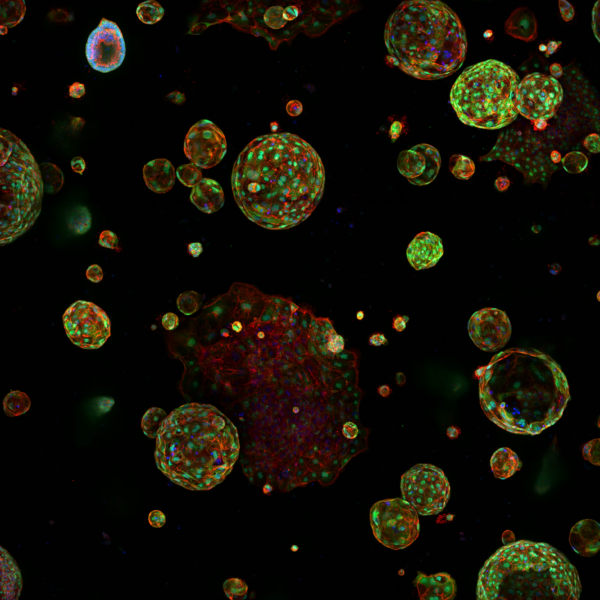
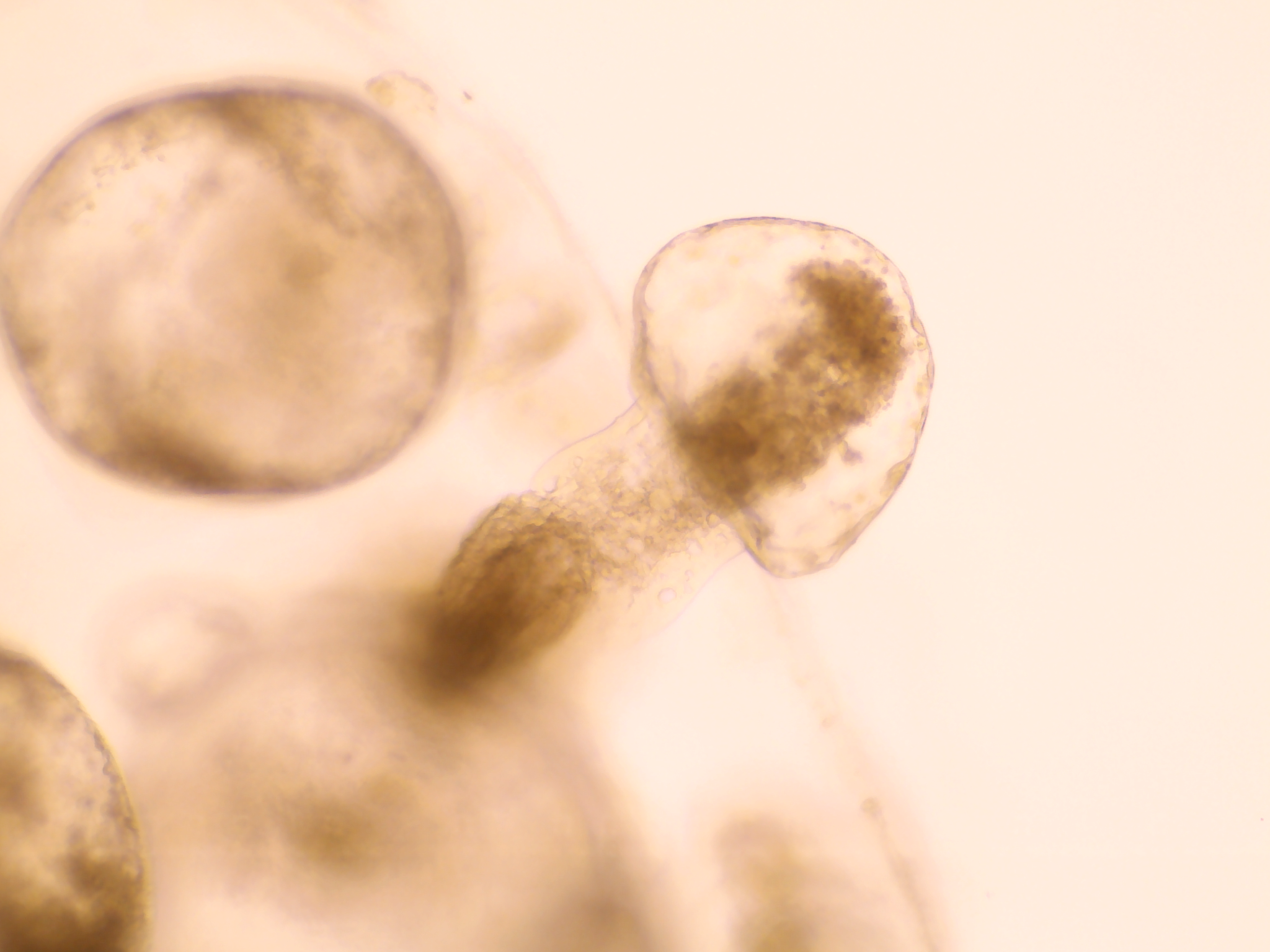
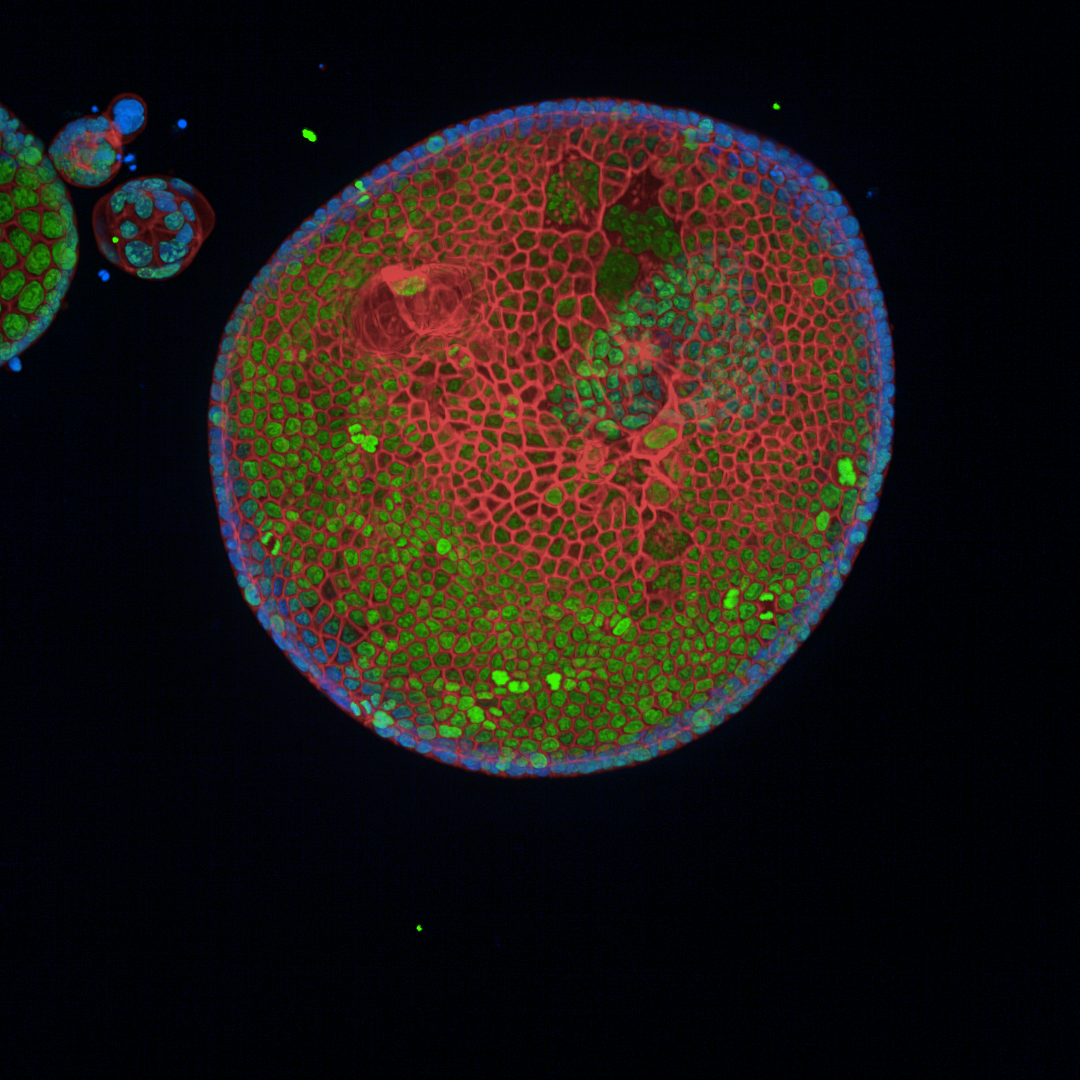
Reducing the stiffness of the PDAC organoid environment results in cells creating ductal-like structures as they develop from original more firm area into the softer area.
Featured Publications
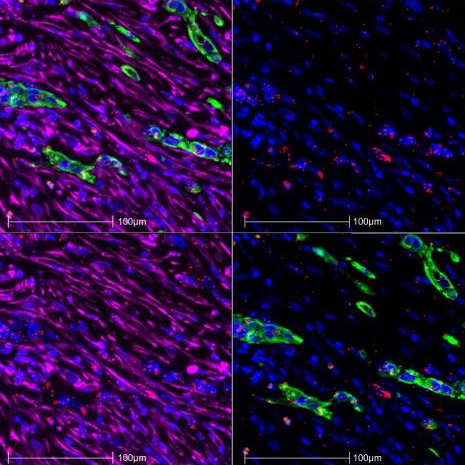
Heterocellular OSM-OSMR signalling reprograms fibroblasts to promote pancreatic cancer growth and metastasis
17th December 2021
Pancreatic ductal adenocarcinoma (PDA) is a lethal malignancy with a complex microenvironment. Here, we describe how heterocellular Oncostatin M (OSM) – Oncostatin M Receptor (OSMR) signalling reprograms fibroblasts, regulates tumour growth and metastasis.
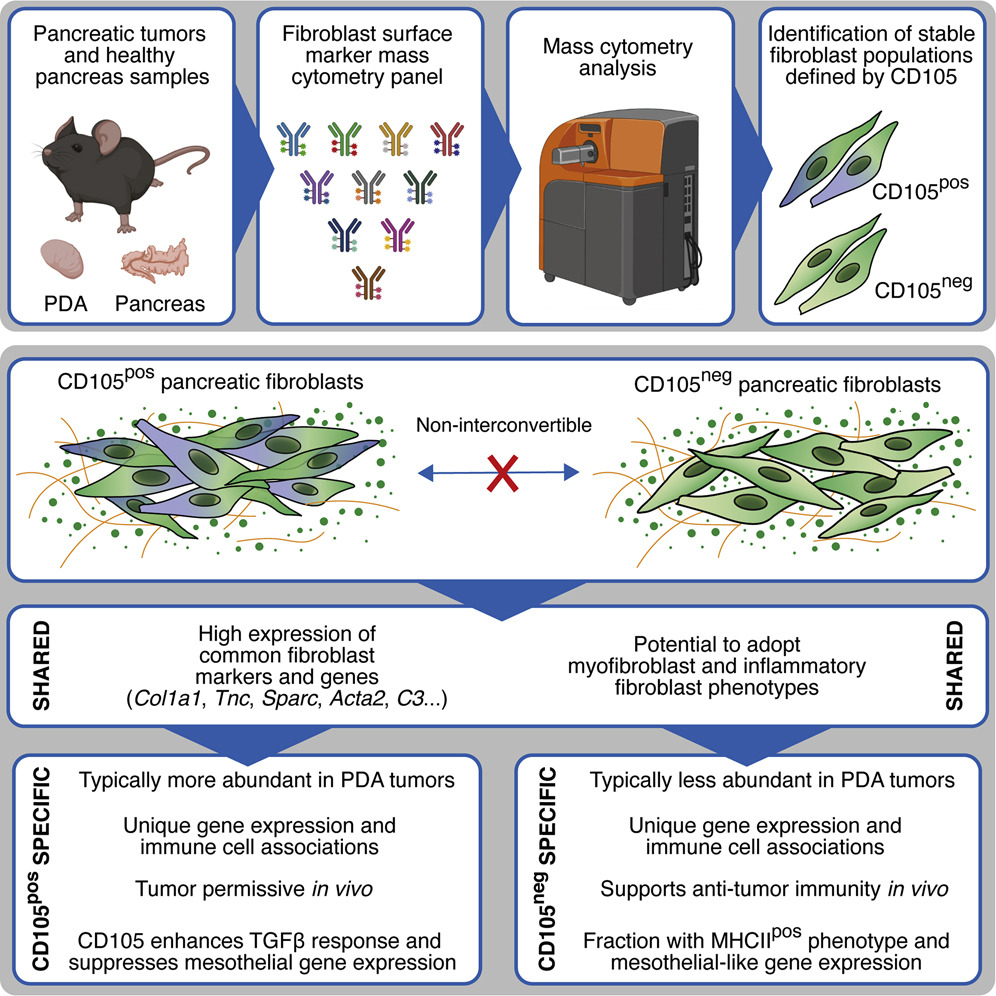
Single-cell analysis defines a pancreatic fibroblast lineage that supports anti-tumor immunity.
13th September 2021
Hutton et al. use mass cytometry to chart stromal cells and describe mesenchymal states and lineages in pancreatic ductal adenocarcinoma. CD105 (Eng) expression distinguishes two pancreatic fibroblast lineages with distinct functions. CD105pos fibroblasts are tumour permissive, whereas CD105neg fibroblasts suppress tumour growth in a manner dependent on adaptive immunity.
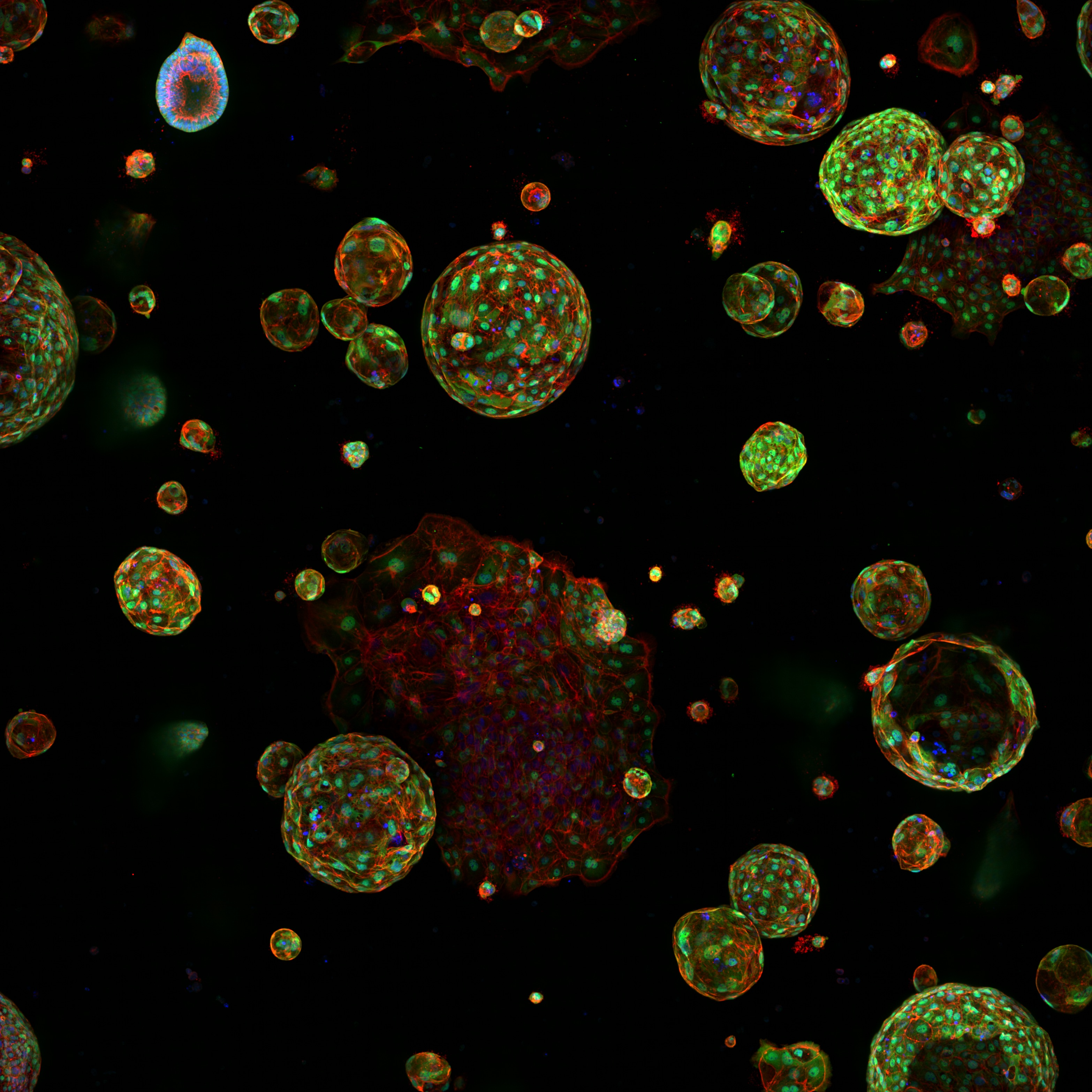
A microenvironment-inspired synthetic three-dimensional model for pancreatic ductal adenocarcinoma organoids
13th September 2021
Below et al. describe a fully synthetic hydrogel extracellular matrix designed to elicit key phenotypic traits of the pancreatic environment in culture.
Stiffness in Pancreatic Cancer
Reducing the stiffness of the PDAC organoid environment results in cells creating ductal-like structures as they develop from original more firm area into the softer area.
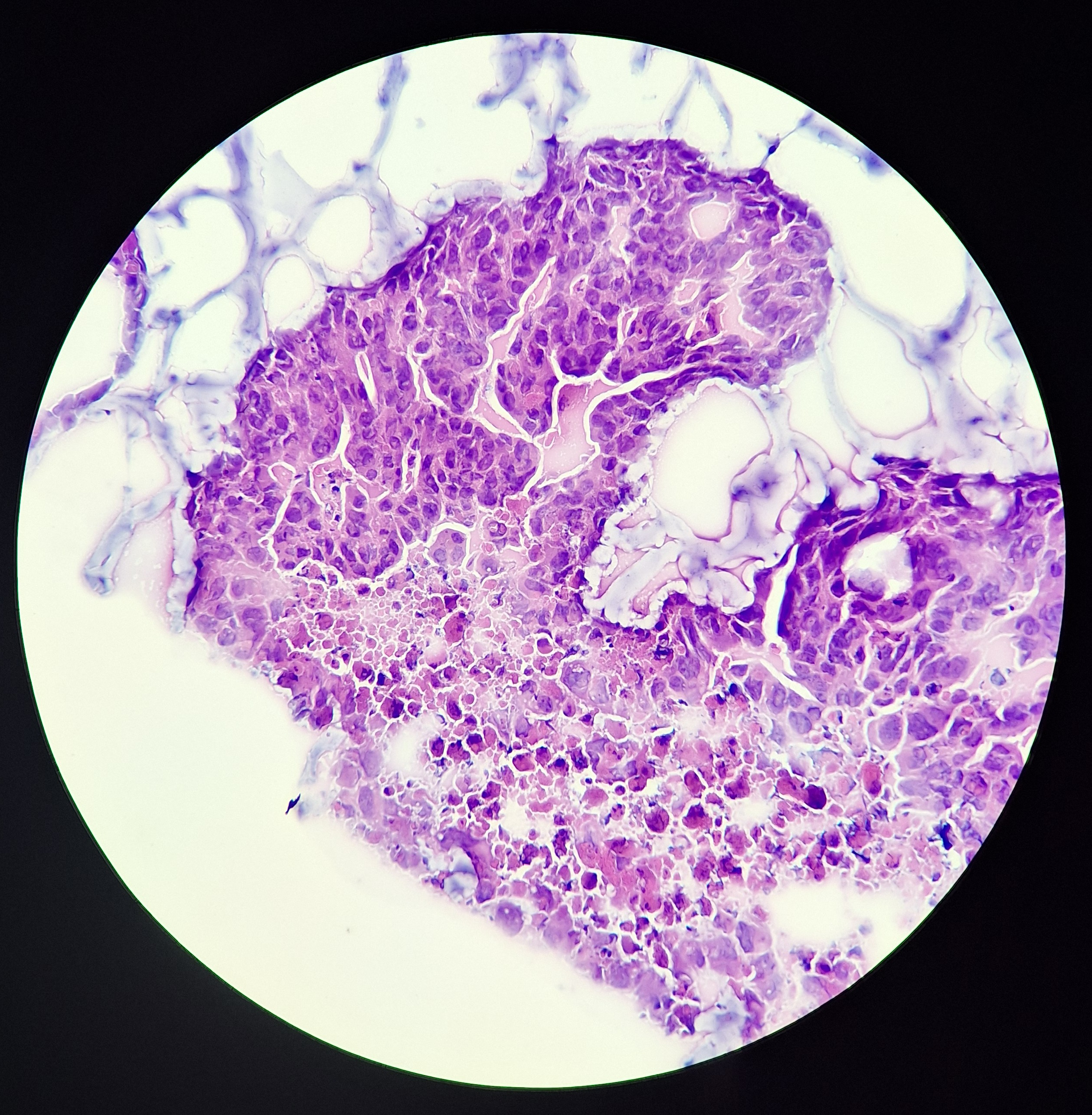
We now know that tumours are a complex mix of genetically diverse cancer cells and multiple types of healthy cells, all communicating with each other via an intricate web of interactions. Untangling this web, and decoding individual signals, is vital to identify which of the multitude of communications are most important for controlling tumour growth and spread. We have identified a key role played by the most commonly mutated gene in cancer in communicating with healthy cells. Blocking its effects could be an effective cancer treatment.
Claus Jorgensen
Senior Group Leader
Meet the Systems Oncology Team
It’s a great pleasure to introduce the Systems Oncology group, who are a joy to work with as they are always eager to support each other and welcome new members to the team. They are dedicated to improving outcomes for cancer patients through working collaboratively on innovative research – and having fun at the same time.

Deputy Director & Senior Group Leader
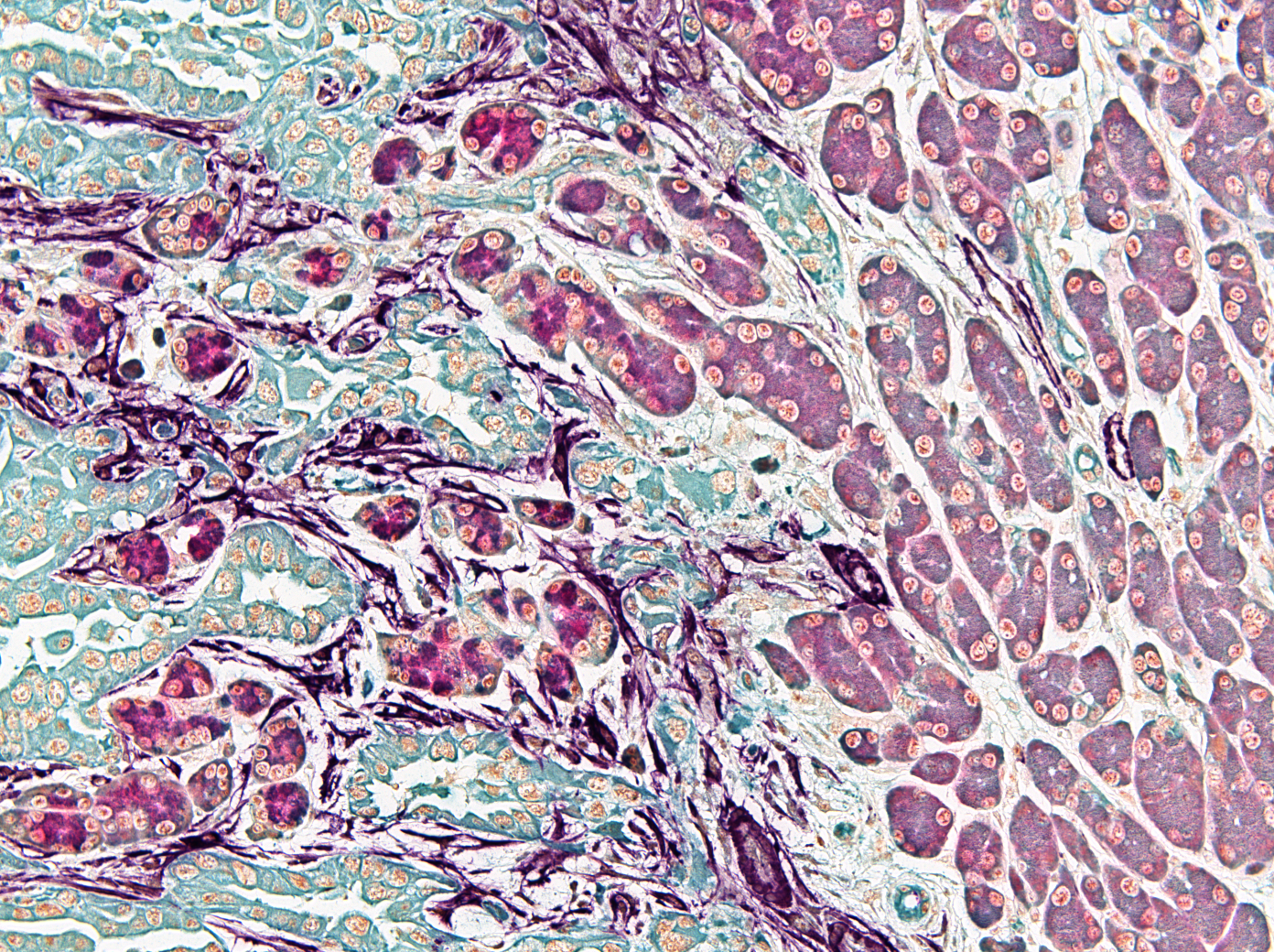
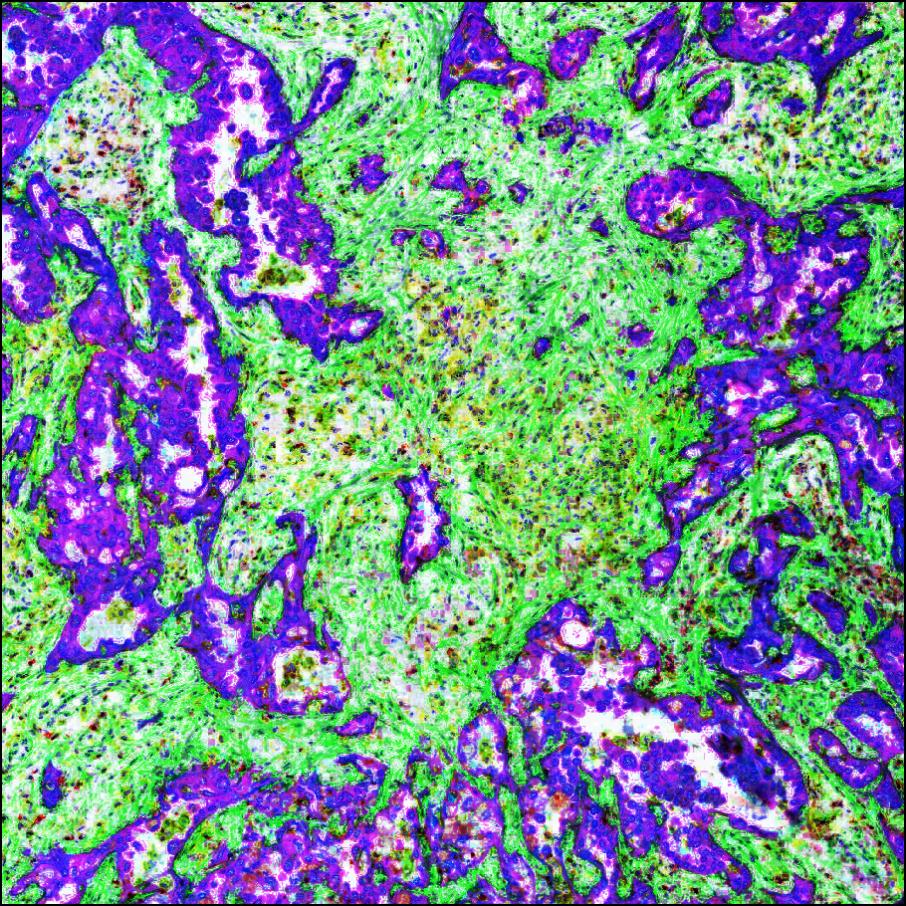
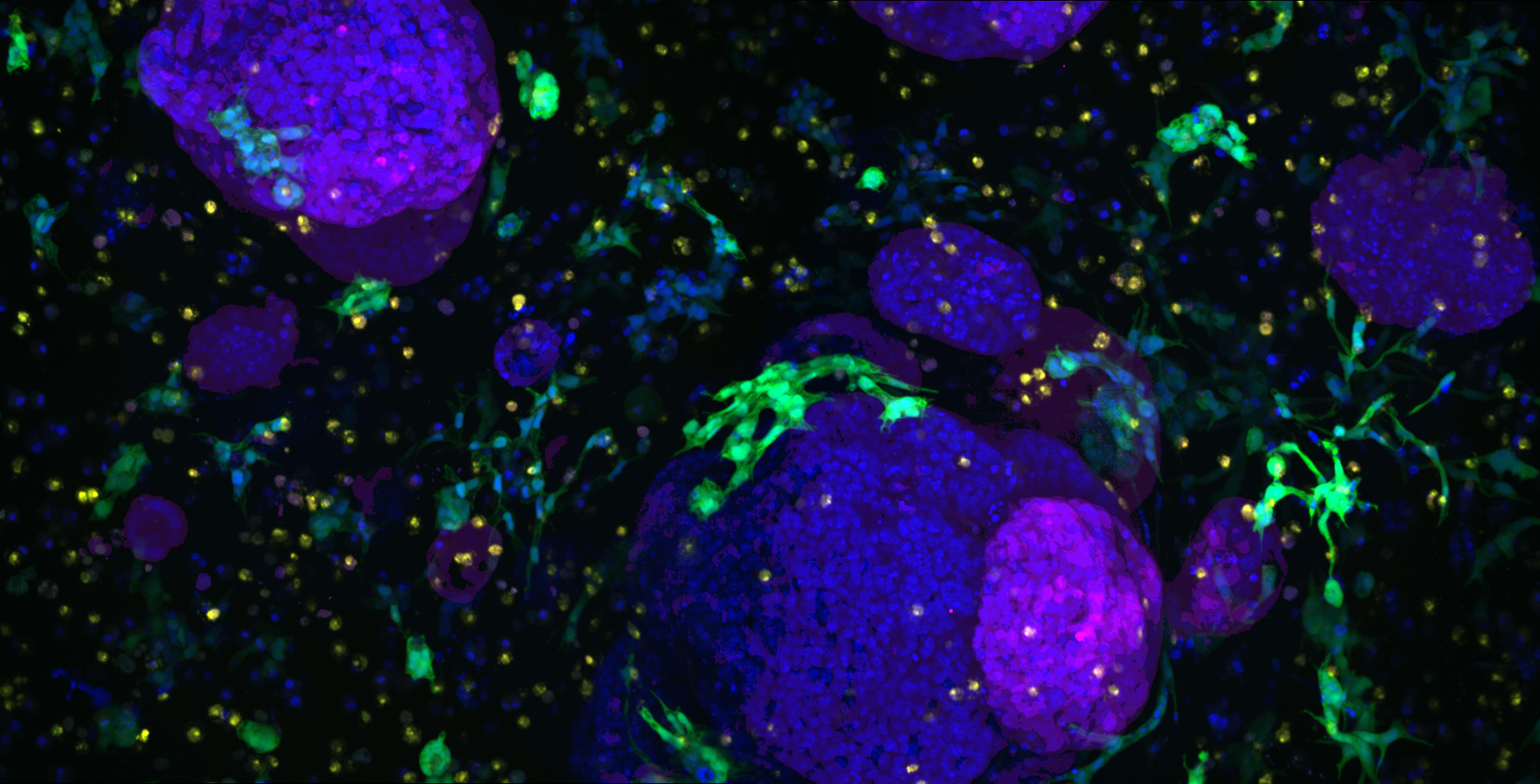
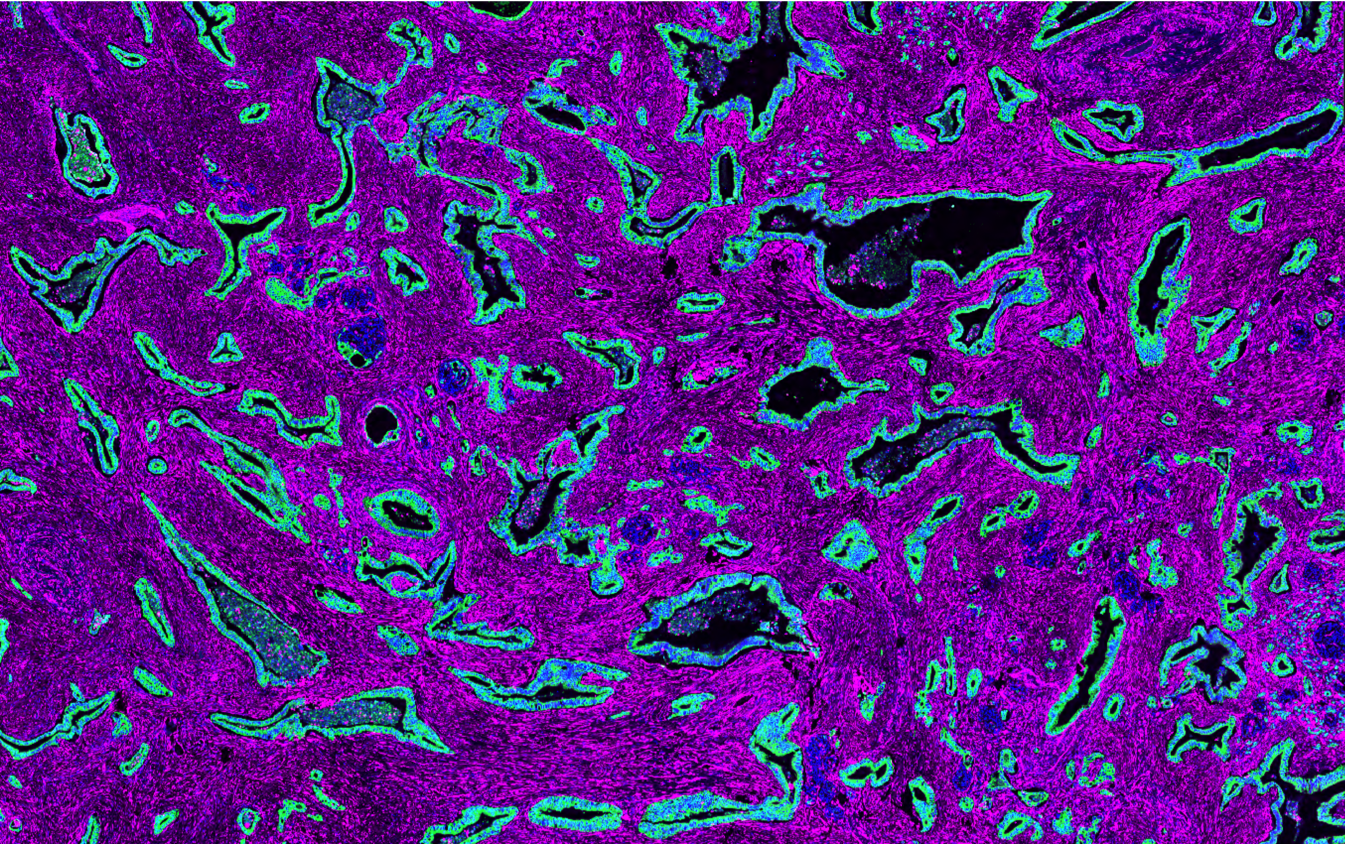
Some of our research images
FAQS
While we are not involved in treating patients, we collaborate with clinicians and the MCRC Biobank to use patient samples in our work. We create 3D tumour models based on patient tissue grown in the laboratory, making these models highly relevant tools to study the growth of pancreatic cancer, the signals sent between cancer and host cells, and for investigating new treatment options.
We work with research partners across the University of Manchester, around the UK and internationally. Get in touch with us using the contact form, we’re always open to new ideas and collaborations.
We are not currently advertising for a PhD student, but please see the student pages for information about our PhD programme. If you are interested in our work, please feel free to get in touch directly using the contact form at any time.
All publications
https://www.nature.com/articles/s41467-026-69964-2
Disruption of tRNA threonylation triggers RIG-I mediated anti-tumour immune response
25 February 2026
Institute Authors (1)
Sylvain Delaunay
Research Group
RNA Dynamics in Cancer
25 February 2026
https://doi.org/10.1038/s44161-025-00740-z
Single-cell profiling reveals three endothelial-to-hematopoietic transitions with divergent isoform expression landscapes
11 November 2025
Institute Authors (6)
Robert Sellers, John Weightman, Wolfgang Breitwieser, Natalia Moncaut, Michael Lie-a-ling, Georges Lacaud
Labs & Facilities
Computational Biology Support, Molecular Biology, Genome Editing and Mouse Models
Research Group
Stem Cell Biology
11 November 2025
https://doi.org/10.1136/jitc-2025-012527
Systemic immunosuppression from ultraviolet radiation exposure inhibits cancer immunotherapy
31 October 2025
Institute Authors (4)
Isabella Mataloni, Antonia Banyard, Garry Ashton, Amaya Virós
Labs & Facilities
Mass and Flow Cytometry, Histology
Research Group
Skin Cancer & Ageing
31 October 2025
https://aacrjournals.org/cancerdiscovery/article/doi/10.1158/2159-8290.CD-24-1224/766638/Glucocorticoids-Unleash-Immune-dependent-Melanoma
Glucocorticoids Unleash Immune-dependent Melanoma Control through Inhibition of the GARP/TGF β Axis
15 October 2025
Institute Authors (12)
Charles Earnshaw, Poppy Dunn, Shih-Chieh Chiang, Maria Koufaki, Massimo Russo, Kimberley Hockenhull, Erin Richardson, Anna Pidoux, Alex Baker, Richard Reeves, Robert Sellers, Sudhakar Sahoo
Labs & Facilities
Computational Biology Support, Visualisation, Irradiation and Analysis
Research Group
Cancer Inflammation and Immunity
15 October 2025
/wp-content/uploads/2025/09/Annual_Report_2024.pdf
2024 Annual Report
23 September 2025
23 September 2025
https://doi.org/10.1182/blood.2024028033
An in vivo barcoded CRISPR-Cas9 screen identifies Ncoa4-mediated ferritinophagy as a dependence in Tet2-deficient hematopoiesis
4 September 2025
Institute Authors (1)
Justin Loke
Research Group
Myeloid Cancer Biology
4 September 2025
Get in touch
Our vision for world leading cancer research in the heart of Manchester
We are a leading cancer research institute within The University of Manchester, spanning the whole spectrum of cancer research – from investigating the molecular and cellular basis of cancer, to translational research and the development of therapeutics.
Our collaborations
Bringing together internationally renowned scientists and clinicians
Scientific Advisory Board
Supported by an international Scientific Advisory Board
Careers that have a lasting impact on cancer research and patient care
We are always on the lookout for talented and motivated people to join us. Whether your background is in biological or chemical sciences, mathematics or finance, computer science or logistics, use the links below to see roles across the Institute in our core facilities, operations teams, research groups, and studentships within our exceptional graduate programme.
















A note from group leader – Claus Jørgensen
My research interest is mainly focused on systems-level analysis of signalling networks in cellular decision processes and how these are deregulated in cancer. I am particularly interested in understanding how aberrations in cellular signalling networks promote tumorigenesis and how the tumour microenvironment affects tumour cell signalling and malignant progression.
In my laboratory we focus on identifying and delineating the regulatory mechanisms whereby tumour cells co-opt stromal cells and how the ‘activated’ stromal cells affect tumour cell behaviour (malignant progression, drug sensitivity and metastasis). To establish the specificities of signalling networks and how they impinge on specific cellular processes, we use Mass Cytometry (CyTOF), CRISPR editing/screening, sequencing and quantitative mass spectrometry to systematically measure signalling on a global level. Through data-integrative approaches, we aim to describe how signals are processed in a cell-specific manner. The long-term goals are to understand how specific signals are processed to promote tumour progression and how blocking these signals can enhance therapeutic response.